Systems Biology
Advertisement
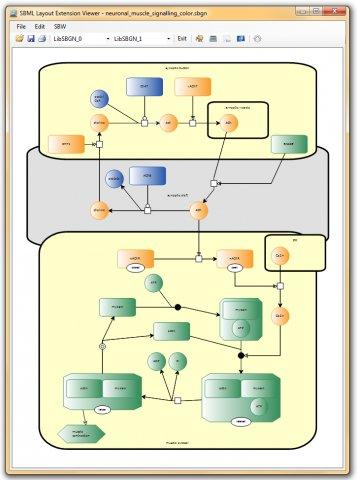
Systems Biology Markup Language (SBML) v.1
The Systems Biology Markup Language (SBML) is an XML-based description language for representing computational models in systems biology.
Advertisement
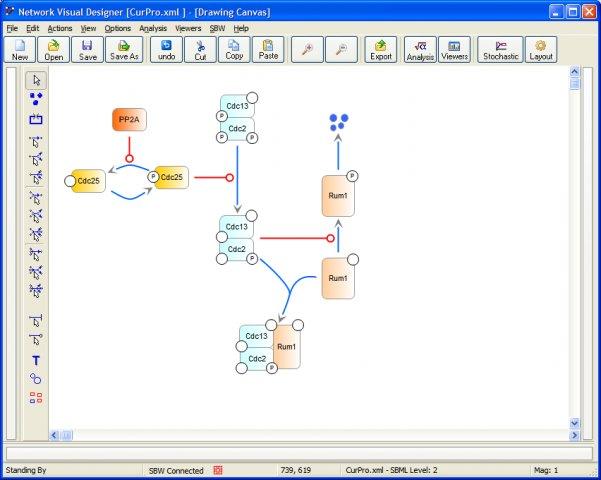
Systems Biology Software Project v.1.0
This site hosts the source code for C++ version of the Broker for SBW, NOM module, advanced simulation suite, analysis applications and model editors.
Electronic Cancer System Studio v.1.0.0.1
A computational model development and simulation environment for cancer systems biology.
Gepasi v.3 3
Gepasi is a software package for modeling biochemical systems.
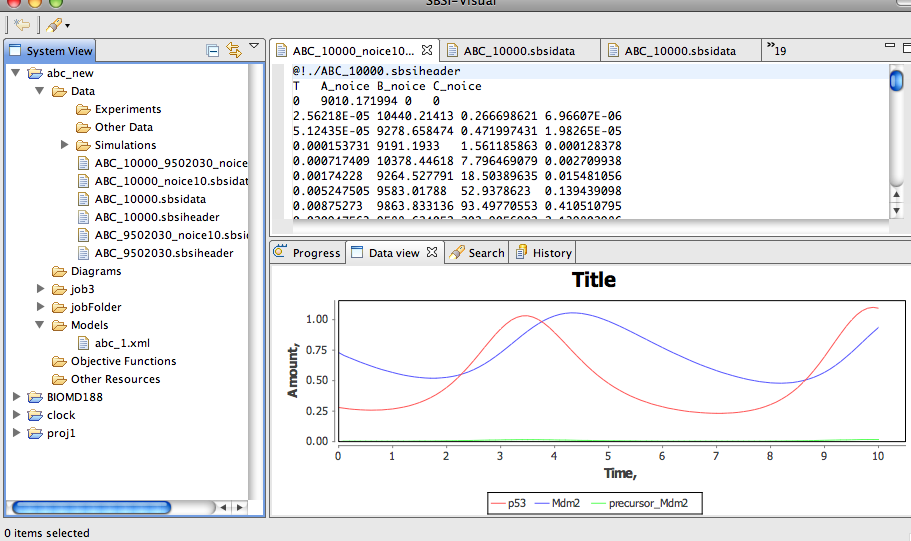
SBSI Visual 1.3.5 Beta v.1.0
SBSI or Systems Biology Software Infrastructure is a handy suite of software tools specially designed for systems biology.
CellDesigner v.4.1.0.6
CellDesigner is a structured diagram editor for drawing gene-regulatory and biochemical networks.
Pathway Tools v.15.0
Pathway Tools is a comprehensive symbolic systems biology software system that supports several use cases in bioinformatics and systems biology: - Development of organism-specific databases - Scientific Visualization, web publishing - Visual anal
ROC.KIT v.1.2.1
ROC.
SBRML.NET - Tools & Library for SBRML v.1.0
The Systems Biology Results Markup Language is a language describing data.
DAMBE (Data Analysis and Molecular Biology and Evolution) v.5.2.30
DAMBE (Data Analysis and Molecular Biology and Evolution) help you with data analysis and molecular biology and evolution.Here is a short summary of DAMBE functions 1. Sequence alignment * General sequence alignment with nucleotide and amino acid
BioTapestry v.5.0.2
BioTapestry is designed around the concept of a developmental network model, and is intended to deal with large scale models with consistency and clarity. It is capable of representing systems that exhibit increasing complexity over time,